|
3/16/2023 0 Comments Macvector protein toolbox 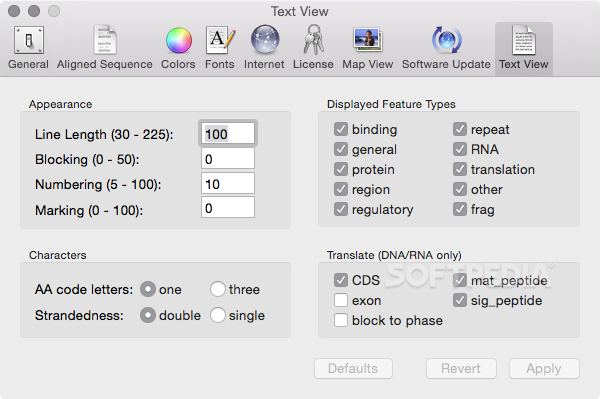
Restriction enzyme‐dependent vectors typically incorporate rare cutting sites or homing endonuclease recognition sites (such as pSAT and pPZP‐RCS) for multigene assembly via a protracted, low‐efficiency cloning process (Goderis et al., 2002 Tzfira et al., 2005). To date, several plant transformation vector systems for multiple FFP expression units have been reported (Patron, 2014 Zhu et al., 2020a, 2020b). Single‐plasmid transformation systems are, therefore, more desirable and require combination of two or more target expression units. The efficiency of such analyses can be highly variable due to gene dosage. FTPL and/or BiFC analyses, however, often require the co‐transformation of two or more plasmids (French et al., 2008 Waadt and Kudla, 2008). Moreover, many Gateway cloning‐based vectors (such as pGWBs, pEarleyGate and pDESTs) are widely utilized in plant research to conveniently transfer DNA fragments between vectors without the need for restriction enzyme digestion (Earley et al., 2006 Gehl et al., 2009 Karimi et al., 2007 Nakagawa et al., 2007).įor most of these applications, the basic designs of the vectors are suitable for manipulation of single gene. A set of ligation‐independent cloning‐compatible vectors (pPLVs) that allow multiple fragments to be assembled in a single reaction were also developed for FTPL analysis in plants (De Rybel et al., 2011). Improved T‐A cloning vectors (such as pXs/pCXs and pUC‐35s‐FPs/pGreen‐Ubi‐FPs) that enable the cloning of PCR fragments are generated for FTPL assays (Chen et al., 2009 Wang et al., 2013). The first vectors developed for FTPL (such as pGDs and pSAT) and BiFC (such as pSYs and pSATNs) assays depended on traditional cloning methods using restriction enzyme digestion and ligation (Bracha‐Drori et al., 2004 Citovsky et al., 2006 Goodin et al., 2002 Tzfira et al., 2005). Over the past decades, many specialized vectors have been developed for the transient or stable expression of fluorescent fusion proteins (FFPs) in plant cells (Blatt and Grefen, 2014). 
The first step in such analyses is the preparation of constructs expressing target fusion proteins. With the development of fluorescent proteins and high‐resolution microscopes, fluorescent tagging protein localization (FTPL) and bimolecular fluorescence complementation (BiFC) technologies have become widely used for PSL and PPI analyses, providing direct visualization in living cells (Dixit et al., 2006 Tanz et al., 2013). This modular UNS‐guided GA‐mediated AioFFP vector toolkit is cost‐effective, easy to use and will promote functional genomics research in plants.ĭefining protein subcellular localization (PSL) and protein–protein interaction (PPI) networks are crucial for studying plant gene functions and cellular processes in the post‐genomics era (Cui et al., 2016 Miller et al., 2015). In addition, we performed a high‐throughput assessment of the accurate subcellular localizations of an uncharacterized rice CBSX protein subfamily. We validated the AioFFP system by testing genes encoding proteins known to be functional in FTPL and BiFC assays. This system also enables integration of organelle marker genes or fluorescently fused target gene expression units into a single transient expression plasmid or binary vector. In brief, this system enables convenient cloning of a target gene into various FFP vectors or the insertion of two or more target genes into the same FFP vector in a single‐tube GA reaction. This toolbox uses Gibson assembly (GA) and incorporates multiple unique nucleotide sequences (UNSs) to facilitate efficient gene cloning. 
Here, to address these needs, we developed an efficient, modular all‐in‐one (Aio) FFP (AioFFP) vector toolbox, including a set of fluorescently labelled organelle markers, FTPL and BiFC plasmids and associated binary vectors. Functional genomics applications using FFPs such as a gene family studies also often require the generation of multiple plasmids. The efficiency of fluorescent fusion protein (FFP) expression analyses is typically impaired when the FFP genes are co‐transformed on separate plasmids compared to when all are cloned and transformed in a single vector. Fluorescent tagging protein localization (FTPL) and bimolecular fluorescence complementation (BiFC) are popular tools for in vivo analyses of the subcellular localizations of proteins and protein–protein interactions in plant cells. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
